plot_feature(input_data,
'age',
xlabel = 'blood serum measurement',
ylabel = 'disease progression')
from helpers_new import *
from sklearn.datasets import load_diabetes
import matplotlib as mpl
from tqdm.notebook import tqdm_notebook as tq
import time
%matplotlib inline
plt.style.use('dark_blue_greens.mplstyle')
css_styling()
| | Top | SciKit-Learn Way | SKTime Way | Multivariate | Panel Data | SKLearn & SKTime | Univariate Forecasting | Advanced Workflow | Forecasting with Exogeneous | Building a Forecaster | Time Series Classification | Time Series Regression | | ||
|---|---|---|
| The SciKit-Learn Way |
| Diabetes Dataset (documentation) |
|---|
Ten baseline variables, age, sex, body mass index, average blood pressure, and six blood serum measurements were obtained for each of n = 442 diabetes patients, as well as the response of interest, a quantitative measure of disease progression one year after baseline.
diabetes = load_diabetes()
input_data = diabetes['data']
target_data = diabetes['target']
input_df = pd.DataFrame(input_data, columns = diabetes['feature_names'])
target_series = pd.Series(target_data)
| Inputs and Targets |
|---|
head_tail_vert(input_df, 5, 'diabetes: input_data')
| diabetes: input_data: head(5) |
| age | sex | bmi | bp | s1 | s2 | s3 | s4 | s5 | s6 | |
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0.04 | 0.05 | 0.06 | 0.02 | -0.04 | -0.03 | -0.04 | -0.00 | 0.02 | -0.02 |
| 1 | -0.00 | -0.04 | -0.05 | -0.03 | -0.01 | -0.02 | 0.07 | -0.04 | -0.07 | -0.09 |
| 2 | 0.09 | 0.05 | 0.04 | -0.01 | -0.05 | -0.03 | -0.03 | -0.00 | 0.00 | -0.03 |
| 3 | -0.09 | -0.04 | -0.01 | -0.04 | 0.01 | 0.02 | -0.04 | 0.03 | 0.02 | -0.01 |
| 4 | 0.01 | -0.04 | -0.04 | 0.02 | 0.00 | 0.02 | 0.01 | -0.00 | -0.03 | -0.05 |
| diabetes: input_data: tail(5) |
| age | sex | bmi | bp | s1 | s2 | s3 | s4 | s5 | s6 | |
|---|---|---|---|---|---|---|---|---|---|---|
| 437 | 0.04 | 0.05 | 0.02 | 0.06 | -0.01 | -0.00 | -0.03 | -0.00 | 0.03 | 0.01 |
| 438 | -0.01 | 0.05 | -0.02 | -0.07 | 0.05 | 0.08 | -0.03 | 0.03 | -0.02 | 0.04 |
| 439 | 0.04 | 0.05 | -0.02 | 0.02 | -0.04 | -0.01 | -0.02 | -0.01 | -0.05 | 0.02 |
| 440 | -0.05 | -0.04 | 0.04 | 0.00 | 0.02 | 0.02 | -0.03 | 0.03 | 0.04 | -0.03 |
| 441 | -0.05 | -0.04 | -0.07 | -0.08 | 0.08 | 0.03 | 0.17 | -0.04 | -0.00 | 0.00 |
head_tail_horz(target_series, 5, "diabetes: target_data")
| diabetes: target_data |
| 0 | |
|---|---|
| 0 | 151.00 |
| 1 | 75.00 |
| 2 | 141.00 |
| 3 | 206.00 |
| 4 | 135.00 |
| 0 | |
|---|---|
| 437 | 178.00 |
| 438 | 104.00 |
| 439 | 132.00 |
| 440 | 220.00 |
| 441 | 57.00 |
| Initial Visualization |
|---|
cols = list(input_df.columns)
def plot_feature(df,
column,
color = None,
title = None,
xlabel = None,
ylabel = None):
if color:
color = color
else:
color = 'C0'
cols = list(input_df.columns)
fig, ax = plt.subplots(1)
col = cols.index(column)
ax.scatter(input_data[:,col], target_data, color = color)
ax.set_title(f'{column} vs. {ylabel}')
ax.set(
xlabel = f'{xlabel}: {diabetes["feature_names"][col]}',
ylabel = f'{ylabel}');
| Visualization: age vs disease progression |
|---|
plot_feature(input_data,
'age',
xlabel = 'blood serum measurement',
ylabel = 'disease progression')
| Visualization: bmi vs disease progression |
|---|
plot_feature(input_data,
'bmi',
xlabel = 'blood serum measurement',
ylabel = 'disease progression',
color = 'C1')
| Visualization: bp vs disease progression |
|---|
plot_feature(input_data,
'bp',
xlabel = 'blood serum measurement',
ylabel = 'disease progression',
color = 'C2')
| Workflow with SciKit-Learn |
|---|
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split
train_in, test_in, train_out, test_out = train_test_split(input_df, target_series)
pretty(f'train_in.shape: {train_in.shape}')
pretty(f'train_out.shape: {train_out.shape}')
pretty(f'test_in.shape: {test_in.shape}')
pretty(f'test_out.shape: {test_out.shape}')
| train_in.shape: (331, 10) |
| train_out.shape: (331,) |
| test_in.shape: (111, 10) |
| test_out.shape: (111,) |
| Model |
|---|
classifier = RandomForestRegressor()
classifier.fit(train_in, train_out)
RandomForestRegressor()In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook.
RandomForestRegressor()
predictions = classifier.predict(test_in)
mean_squared_error(test_out, predictions)
3599.5111306306303
| Modular Model Building & Pipelines with SciKit-Learn |
|---|
from sklearn.neighbors import KNeighborsRegressor
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
pipeline = make_pipeline(StandardScaler(), KNeighborsRegressor())
pipeline
Pipeline(steps=[('standardscaler', StandardScaler()),
('kneighborsregressor', KNeighborsRegressor())])In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. Pipeline(steps=[('standardscaler', StandardScaler()),
('kneighborsregressor', KNeighborsRegressor())])StandardScaler()
KNeighborsRegressor()
pipeline.fit(train_in, train_out)
Pipeline(steps=[('standardscaler', StandardScaler()),
('kneighborsregressor', KNeighborsRegressor())])In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. Pipeline(steps=[('standardscaler', StandardScaler()),
('kneighborsregressor', KNeighborsRegressor())])StandardScaler()
KNeighborsRegressor()
predictions = pipeline.predict(test_in)
mean_squared_error(test_out, predictions)
3670.1308108108105
| Summary |
|---|
| | Top | SciKit-Learn Way | SKTime Way | Multivariate | Panel Data | SKLearn & SKTime | Univariate Forecasting | Advanced Workflow | Forecasting with Exogeneous | Building a Forecaster | Time Series Classification | Time Series Regression | | ||
|---|---|---|
| The SKTime Way |
| Lynx Dataset (documentation) |
|---|
The annual numbers of lynx trappings for 1821–1934 in Canada. This time-series records the number of skins of predators (lynx) that were collected over several years by the Hudson’s Bay Company. The dataset was taken from Brockwell & Davis (1991) and appears to be the series considered by Campbell & Walker (1977).
Dimensionality: univariate Series length: 114 Frequency: Yearly Number of cases: 1
This data shows aperiodic, cyclical patterns, as opposed to periodic, seasonal patterns.
from sktime.datasets import load_lynx
from sktime.utils.plotting import plot_series
lynx = load_lynx()
plot_series(lynx, title = 'Plotting a Univariate Time Series', colors = ['C4']);
| | Top | SciKit-Learn Way | SKTime Way | Multivariate | Panel Data | SKLearn & SKTime | Univariate Forecasting | Advanced Workflow | Forecasting with Exogeneous | Building a Forecaster | Time Series Classification | Time Series Regression | | ||
|---|---|---|
| Multivariate Data |
| Longley Dataset (documentation) |
|---|
This mulitvariate time series dataset contains various US macroeconomic variables from 1947 to 1962 that are known to be highly collinear.
Dimensionality: multivariate, 6 Series length: 16 Frequency: Yearly Number of cases: 1
Variable description:
TOTEMP - Total employment GNPDEFL - Gross national product deflator GNP - Gross national product UNEMP - Number of unemployed ARMED - Size of armed forces POP - Population
from sktime.datasets import load_longley
targets, inputs = load_longley()
head_tail_horz(inputs, 5, 'Inputs')
| Inputs |
| GNPDEFL | GNP | UNEMP | ARMED | POP | |
|---|---|---|---|---|---|
| Period | |||||
| 1947 | 83.00 | 234,289.00 | 2,356.00 | 1,590.00 | 107,608.00 |
| 1948 | 88.50 | 259,426.00 | 2,325.00 | 1,456.00 | 108,632.00 |
| 1949 | 88.20 | 258,054.00 | 3,682.00 | 1,616.00 | 109,773.00 |
| 1950 | 89.50 | 284,599.00 | 3,351.00 | 1,650.00 | 110,929.00 |
| 1951 | 96.20 | 328,975.00 | 2,099.00 | 3,099.00 | 112,075.00 |
| GNPDEFL | GNP | UNEMP | ARMED | POP | |
|---|---|---|---|---|---|
| Period | |||||
| 1958 | 110.80 | 444,546.00 | 4,681.00 | 2,637.00 | 121,950.00 |
| 1959 | 112.60 | 482,704.00 | 3,813.00 | 2,552.00 | 123,366.00 |
| 1960 | 114.20 | 502,601.00 | 3,931.00 | 2,514.00 | 125,368.00 |
| 1961 | 115.70 | 518,173.00 | 4,806.00 | 2,572.00 | 127,852.00 |
| 1962 | 116.90 | 554,894.00 | 4,007.00 | 2,827.00 | 130,081.00 |
head_tail_horz(targets, 5, 'Targets')
| Targets |
| TOTEMP | |
|---|---|
| Period | |
| 1947 | 60,323.00 |
| 1948 | 61,122.00 |
| 1949 | 60,171.00 |
| 1950 | 61,187.00 |
| 1951 | 63,221.00 |
| TOTEMP | |
|---|---|
| Period | |
| 1958 | 66,513.00 |
| 1959 | 68,655.00 |
| 1960 | 69,564.00 |
| 1961 | 69,331.00 |
| 1962 | 70,551.00 |
| Visualization: multiple input series |
|---|
plot_series(targets, colors = ["C0"])
for idx, column in enumerate(inputs.columns[:2]):
current = inputs[column]
plot_series(current, colors = ["C" + str(idx+1)])
| Arrowhead Dataset (documentation) |
|---|
Dimensionality: univariate Series length: 251 Train cases: 36 Test cases: 175 Number of classes: 3
The arrowhead data consists of outlines of the images of arrowheads. The shapes of the projectile points are converted into a time series using the angle-based method. The classification of projectile points is an important topic in anthropology. The classes are based on shape distinctions such as the presence and location of a notch in the arrow. The problem in the repository is a length normalised version of that used in Ye09shapelets. The three classes are called “Avonlea”, “Clovis” and “Mix”.”
import matplotlib.pyplot as plt
from sktime.datasets import load_arrow_head
from sktime.datatypes import convert
inputs, targets = load_arrow_head(return_X_y = True)
head_tail_horz(inputs, 5, 'input data')
| input data |
| dim_0 | |
|---|---|
| 0 | 0 -1.96 1 -1.96 2 -1.96 3 -1.94 4 -1.90 ... 246 -1.84 247 -1.88 248 -1.91 249 -1.92 250 -1.91 Length: 251, dtype: float64 |
| 1 | 0 -1.77 1 -1.77 2 -1.78 3 -1.73 4 -1.70 ... 246 -1.64 247 -1.68 248 -1.73 249 -1.78 250 -1.79 Length: 251, dtype: float64 |
| 2 | 0 -1.87 1 -1.84 2 -1.84 3 -1.81 4 -1.76 ... 246 -1.83 247 -1.88 248 -1.86 249 -1.86 250 -1.85 Length: 251, dtype: float64 |
| 3 | 0 -2.07 1 -2.07 2 -2.04 3 -2.04 4 -1.96 ... 246 -1.95 247 -2.01 248 -2.03 249 -2.07 250 -2.08 Length: 251, dtype: float64 |
| 4 | 0 -1.75 1 -1.74 2 -1.72 3 -1.70 4 -1.68 ... 246 -1.72 247 -1.74 248 -1.74 249 -1.76 250 -1.76 Length: 251, dtype: float64 |
| dim_0 | |
|---|---|
| 206 | 0 -1.63 1 -1.62 2 -1.63 3 -1.61 4 -1.57 ... 246 -1.57 247 -1.60 248 -1.62 249 -1.62 250 -1.62 Length: 251, dtype: float64 |
| 207 | 0 -1.66 1 -1.66 2 -1.63 3 -1.61 4 -1.59 ... 246 -1.68 247 -1.67 248 -1.67 249 -1.68 250 -1.68 Length: 251, dtype: float64 |
| 208 | 0 -1.60 1 -1.59 2 -1.58 3 -1.56 4 -1.53 ... 246 -1.58 247 -1.59 248 -1.60 249 -1.61 250 -1.61 Length: 251, dtype: float64 |
| 209 | 0 -1.74 1 -1.74 2 -1.73 3 -1.72 4 -1.70 ... 246 -1.64 247 -1.67 248 -1.70 249 -1.71 250 -1.73 Length: 251, dtype: float64 |
| 210 | 0 -1.63 1 -1.63 2 -1.62 3 -1.61 4 -1.58 ... 246 -1.51 247 -1.55 248 -1.58 249 -1.60 250 -1.62 Length: 251, dtype: float64 |
head_tail_horz(targets, 5, 'target data')
| target data |
| 0 | |
|---|---|
| 0 | 0 |
| 1 | 1 |
| 2 | 2 |
| 3 | 0 |
| 4 | 1 |
| 0 | |
|---|---|
| 206 | 2 |
| 207 | 2 |
| 208 | 2 |
| 209 | 2 |
| 210 | 2 |
pretty(inputs.shape, 'inputs.shape before conversion')
| inputs.shape before conversion |
| (211, 1) |
inputs = convert(inputs, from_type = 'nested_univ', to_type = 'numpy3D')
pretty(inputs.shape, 'inputs.shape after conversion')
| inputs.shape after conversion |
| (211, 1, 251) |
labels, counts = np.unique(targets, return_counts = True)
| Visualization: multiple input samples |
|---|
fig, ax = plt.subplots(1, figsize = plt.figaspect(0.25))
for label in labels:
ax.plot(inputs[targets == label, 0, :][0], label = f'class {label}');
ax.set(ylabel = 'Scaled Distance from Midpoint', xlabel = 'Index');
ax.set_title('Panel Data: Each line represents a different, independent sample');
labels, counts = np.unique(targets, return_counts = True)
fig, ax = plt.subplots(1, figsize = plt.figaspect(0.25))
for label in labels:
for idx in range(3):
ax.plot(inputs[targets == label, 0, :][idx], label = f'class {label}');
ax.set(ylabel = 'Scaled Distance from Midpoint', xlabel = 'Index');
ax.set_title('Panel Data: Each line represents a different, independent sample');
| | Top | SciKit-Learn Way | SKTime Way | Multivariate | Panel Data | SKLearn & SKTime | Univariate Forecasting | Advanced Workflow | Forecasting with Exogeneous | Building a Forecaster | Time Series Classification | Time Series Regression | | ||
|---|---|---|
| SKLearn + SKTime: Multiple Learning Tasks |
| Reduction: From one learning task to another |
|---|
**Overview**
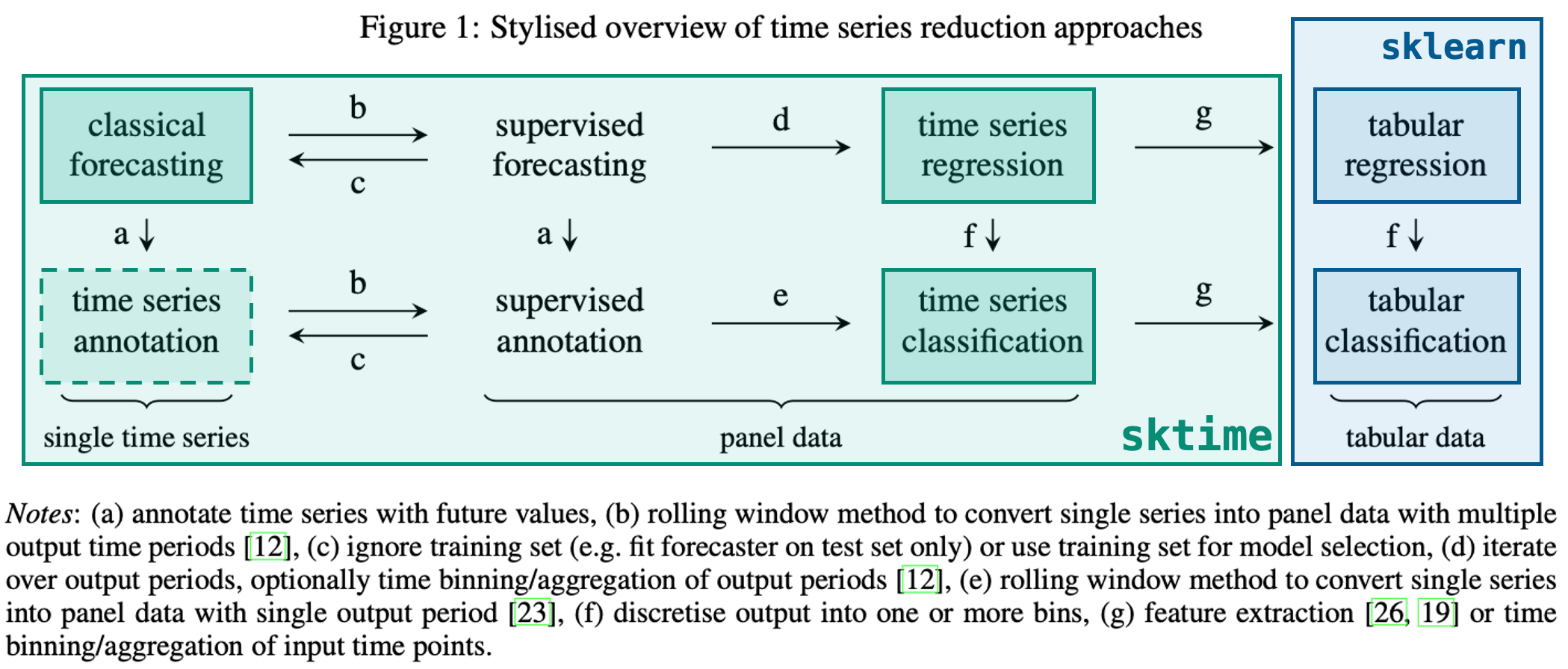
**Example: From forecasting to regression**
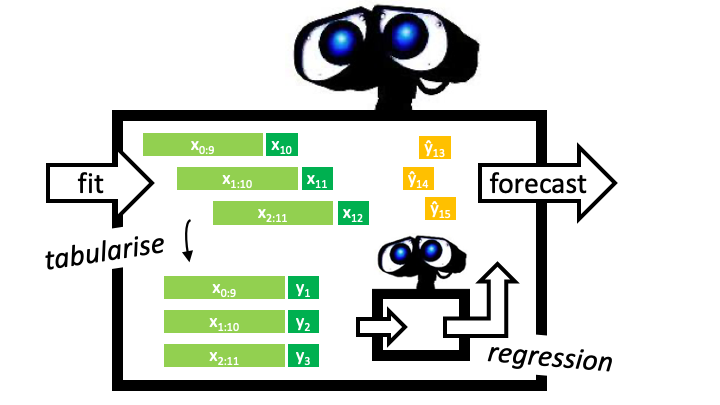
| Creating a unified framework |
|---|

| What's a framework? |
|---|
Check out our glossary of common terms:
A collection of related and reusable software design templates that practitioners can copy and fill in. Frameworks emphasize design reuse. They capture common software design decisions within a given application domain and distill them into reusable design templates. This reduces the design decision they must take, allowing them to focus on application specifics. Not only can practitioners write software faster as a result, but applications will have a similar structure. Frameworks often offer additional functionality like toolboxes. Compare with toolbox and application.
Check out our extension templates!

| | Top | SciKit-Learn Way | SKTime Way | Multivariate | Panel Data | SKLearn & SKTime | Univariate Forecasting | Advanced Workflow | Forecasting with Exogeneous | Building a Forecaster | Time Series Classification | Time Series Regression | | ||
|---|---|---|
| Univariate Forecasting |
| Univariate forecasting |
|---|
In forecasting, we're interested in using past data to make temporal forward predictions. sktime provides common statistical forecasting algorithms and tools for building composite machine learning models.
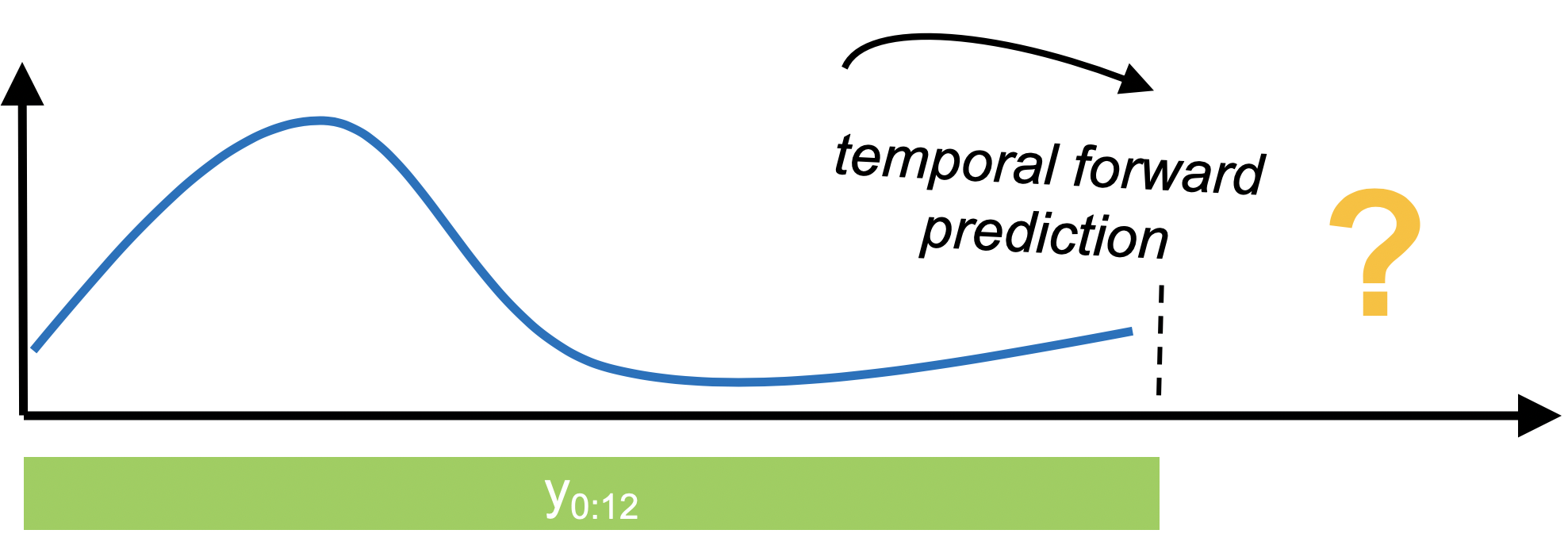
| The basic workflow |
|---|
from warnings import simplefilter
simplefilter(action="ignore", category=RuntimeWarning)
from sktime.datasets import load_shampoo_sales
from sktime.utils.plotting import plot_series
| Data Specification |
|---|
| Shampoo Sales Dataset (documentation) |
|---|
This dataset describes the monthly number of sales of shampoo over a 3 year period. The units are a sales count.
Dimensionality: univariate Series length: 36 Frequency: Monthly Number of cases: 1
shampoo_data = load_shampoo_sales()
plot_series(shampoo_data, title = "Shampoo Sales Time Series",
colors = ['C4']);
| Task specification |
|---|
Next we will define a **forecasting task**
| The Forecasting Horizon |
|---|
When we want to generate forecasts, we need to specify the forecasting horizon and pass that to our forecasting algorithm. We can specify the forecasting horizon as a numpy array of the steps ahead relative to the end of the training series:
ForcastingHorizon()is_relative = False - to specify exact dates. True (default) means that each time point will be relative to the last time point in the dataInit signature:
ForecastingHorizon(
values: Union[int, list, numpy.ndarray, pandas.core.indexes.base.Index] = None,
is_relative: bool = None,
freq=None,
)
Docstring:
Forecasting horizon.
Parameters
----------
values : pd.Index, pd.TimedeltaIndex, np.array, list, pd.Timedelta, or int
Values of forecasting horizon
is_relative : bool, optional (default=None)
- If True, a relative ForecastingHorizon is created:
values are relative to end of training series.
- If False, an absolute ForecastingHorizon is created:
values are absolute.
- if None, the flag is determined automatically:
relative, if values are of supported relative index type
absolute, if not relative and values of supported absolute index type
freq : str, pd.Index, pandas offset, or sktime forecaster, optional (default=None)
object carrying frequency information on values
ignored unless values is without inferrable freqimport numpy as np
horizon = np.arange(6) + 1
pretty(horizon, 'Forecasting Horizon: np.arange(6) + 1 -> Relative')
| Forecasting Horizon: np.arange(6) + 1 -> Relative |
| [1, 2, 3, 4, 5, 6] |
import pandas as pd
from sktime.forecasting.base import ForecastingHorizon
horizon = ForecastingHorizon(
pd.period_range("1993-07", periods=6, freq="M"), is_relative=False
)
pretty(horizon, 'Forecasting Horizon: pd.period_range("1993-07", periods=6, freq="M) -> Absolute')
| Forecasting Horizon: pd.period_range("1993-07", periods=6, freq="M) -> Absolute |
| ForecastingHorizon(['1993-07', '1993-08', '1993-09', '1993-10', '1993-11', '1993-12'], dtype='period[M]', is_relative=False) |
| Converting between absolute and relative forecast horizons |
|---|
to_relative() - cutoff allows you to determin when/where to convert to relative from absolute
Signature: ForecastingHorizon.to_relative(self, cutoff=None) Docstring: Return forecasting horizon values relative to a cutoff.
Parameters
cutoff : pd.Period, pd.Timestamp, int, or pd.Index, optional (default=None)
Cutoff value required to convert a relative forecasting
horizon to an absolute one (and vice versa).
If pd.Index, last/latest value is considered the cutoff
Returns
fh : ForecastingHorizon
Relative representation of forecasting horizon.cutoff = pd.Period("1993-06", freq="M")
pretty(horizon.to_relative(cutoff), 'cutoff = pd.Period("1993-06", freq="M") | horizon.to_relative(cutoff)')
| cutoff = pd.Period("1993-06", freq="M") | horizon.to_relative(cutoff) |
| ForecastingHorizon([1, 2, 3, 4, 5, 6], dtype='int64', is_relative=True) |
| Splitting: temporal_train_test_split() |
|---|
Signature:
temporal_train_test_split(
y: Union[pandas.core.series.Series, pandas.core.frame.DataFrame, numpy.ndarray, pandas.core.indexes.base.Index],
X: Optional[pandas.core.frame.DataFrame] = None,
test_size: Union[int, float, NoneType] = None,
train_size: Union[int, float, NoneType] = None,
fh: Union[int, list, numpy.ndarray, pandas.core.indexes.base.Index, sktime.forecasting.base._fh.ForecastingHorizon, NoneType] = None,
) -> Union[Tuple[pandas.core.series.Series, pandas.core.series.Series], Tuple[pandas.core.series.Series, pandas.core.series.Series, pandas.core.frame.DataFrame, pandas.core.frame.DataFrame]]
Docstring:
Split arrays or matrices into sequential train and test subsets.
Creates train/test splits over endogenous arrays an optional exogenous
arrays.
This is a wrapper of scikit-learn's ``train_test_split`` that
does not shuffle the data.
Parameters
----------
y : pd.Series
Target series
X : pd.DataFrame, optional (default=None)
Exogenous data
test_size : float, int or None, optional (default=None)
If float, should be between 0.0 and 1.0 and represent the proportion
of the dataset to include in the test split. If int, represents the
relative number of test samples. If None, the value is set to the
complement of the train size. If ``train_size`` is also None, it will
be set to 0.25.
train_size : float, int, or None, (default=None)
If float, should be between 0.0 and 1.0 and represent the
proportion of the dataset to include in the train split. If
int, represents the relative number of train samples. If None,
the value is automatically set to the complement of the test size.
fh : ForecastingHorizon
Returns
-------
splitting : tuple, length=2 * len(arrays)
List containing train-test split of `y` and `X` if given.from sktime.forecasting.model_selection import temporal_train_test_split
train_targets, test_targets = temporal_train_test_split(shampoo_data,
fh=horizon)
| Visualization: forecast horizon for train-test split |
|---|
plot_series(train_targets,
test_targets,
labels=["train_targets", "test_targets"],
colors = ['C4', 'C0'],
title = 'Using Forecast Horizon to Train-Test Split');
| Model Specification: NaiveForecaster() |
|---|
**NaiveForecaster** is a forecaster that makes forecasts using simple strategies. Two out of three strategies are robust against NaNs. The NaiveForecaster can also be used for multivariate data and it then applies internally the ColumnEnsembleForecaster, so each column is forecasted with the same strategy.
Internally, this forecaster does the following:
To compute prediction quantiles, we first estimate the standard error of prediction residuals under the assumption of uncorrelated residuals. The forecast variance is then computed by multiplying the residual variance by a constant. This constant is a small-sample bias adjustment and each method (mean, last, drift) have different formulas for computing the constant. These formulas can be found in the Forecasting: Principles and Practice textbook (Table 5.2) [1]_. Lastly, under the assumption that residuals follow a normal distribution, we use the forecast variance and z-scores of a normal distribution to estimate the prediction quantiles.
Parameters
----------
strategy : {"last", "mean", "drift"}, default="last"
Strategy used to make forecasts:
* "last": (robust against NaN values)
forecast the last value in the
training series when sp is 1.
When sp is not 1,
last value of each season
in the last window will be
forecasted for each season.
* "mean": (robust against NaN values)
forecast the mean of last window
of training series when sp is 1.
When sp is not 1, mean of all values
in a season from last window will be
forecasted for each season.
* "drift": (not robust against NaN values)
forecast by fitting a line between the
first and last point of the window and
extrapolating it into the future.
sp : int, or None, default=1
Seasonal periodicity to use in the seasonal forecasting. None=1.
window_length : int or None, default=None
Window length to use in the `mean` strategy. If None, entire training
series will be used.from sktime.forecasting.naive import NaiveForecaster
forecaster = NaiveForecaster(strategy="drift", window_length=10)
| Model Fitting: NaiveForecaster.fit() |
|---|
y - always requiredX - exogenous dataIn an economic model, an exogenous variable is one whose measure is determined outside the model and is imposed on the model, and an exogenous change is a change in an exogenous variable
fh - can pass forecast horizon either in fitting or predictingSignature: forecaster.fit(y, X=None, fh=None)
Docstring:
Fit forecaster to training data.
State change:
Changes state to "fitted".
Writes to self:
Sets self._is_fitted flag to True.
Writes self._y and self._X with `y` and `X`, respectively.
Sets self.cutoff and self._cutoff to last index seen in `y`.
Sets fitted model attributes ending in "_".
Stores fh to self.fh if fh is passed.
Parameters
----------
y : time series in sktime compatible data container format
Time series to which to fit the forecaster.
y can be in one of the following formats:
Series scitype: pd.Series, pd.DataFrame, or np.ndarray (1D or 2D)
for vanilla forecasting, one time series
Panel scitype: pd.DataFrame with 2-level row MultiIndex,
3D np.ndarray, list of Series pd.DataFrame, or nested pd.DataFrame
for global or panel forecasting
Hierarchical scitype: pd.DataFrame with 3 or more level row MultiIndex
for hierarchical forecasting
Number of columns admissible depend on the "scitype:y" tag:
if self.get_tag("scitype:y")=="univariate":
y must have a single column/variable
if self.get_tag("scitype:y")=="multivariate":
y must have 2 or more columns
if self.get_tag("scitype:y")=="both": no restrictions on columns apply
For further details:
on usage, see forecasting tutorial examples/01_forecasting.ipynb
on specification of formats, examples/AA_datatypes_and_datasets.ipynb
fh : int, list, np.array or ForecastingHorizon, optional (default=None)
The forecasting horizon encoding the time stamps to forecast at.
if self.get_tag("requires-fh-in-fit"), must be passed, not optional
X : time series in sktime compatible format, optional (default=None)
Exogeneous time series to fit to
Should be of same scitype (Series, Panel, or Hierarchical) as y
if self.get_tag("X-y-must-have-same-index"), X.index must contain y.index
there are no restrictions on number of columns (unlike for y)forecaster.fit(train_targets)
NaiveForecaster(strategy='drift', window_length=10)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook.
NaiveForecaster(strategy='drift', window_length=10)
| Predicting: NaiveForecaster.predict() |
|---|
Signature: NaiveForecaster.predict(self, fh=None, X=None)
Docstring:
Forecast time series at future horizon.
State required:
Requires state to be "fitted".
Accesses in self:
Fitted model attributes ending in "_".
self.cutoff, self._is_fitted
Writes to self:
Stores fh to self.fh if fh is passed and has not been passed previously.
Parameters
----------
fh : int, list, np.array or ForecastingHorizon, optional (default=None)
The forecasting horizon encoding the time stamps to forecast at.
if has not been passed in fit, must be passed, not optional
X : time series in sktime compatible format, optional (default=None)
Exogeneous time series to fit to
Should be of same scitype (Series, Panel, or Hierarchical) as y in fit
if self.get_tag("X-y-must-have-same-index"), X.index must contain fh.index
there are no restrictions on number of columns (unlike for y)
Returns
-------
y_pred : time series in sktime compatible data container format
Point forecasts at fh, with same index as fh
y_pred has same type as the y that has been passed most recently:
Series, Panel, Hierarchical scitype, same format (see above)predictions = forecaster.predict(horizon)
plot_series(train_targets,
test_targets,
predictions,
labels=['train_targets', 'test_targets', 'predictions'],
colors = ['C4', 'C0', 'C3'],
title = 'Linear Strategy Prediction');
| Evaluation: mean_absolute_percentage_error() |
|---|
from sktime.performance_metrics.forecasting import \
mean_absolute_percentage_error
mean_absolute_percentage_error(test_targets, predictions, symmetric=False)
0.16469764622516225
| Another Example: AutoARIMA |
|---|
AutoARIMA() rather than NaiveForecaster()from sktime.forecasting.arima import AutoARIMA
data = load_shampoo_sales()
training, testing = temporal_train_test_split(data, fh=horizon)
forecaster = AutoARIMA(sp=12, suppress_warnings=True)
forecaster.fit(training)
AutoARIMA(sp=12, suppress_warnings=True)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook.
AutoARIMA(sp=12, suppress_warnings=True)
predictions = forecaster.predict(horizon)
plot_series(training, testing, predictions,
labels=["y_train", "y_test", "y_pred"],
colors = ['C4', 'C0', 'C3'],
title = 'Example: using AutoARIMA as a Model');
| Summary of basic workflow |
|---|
| Forecasters in sktime |
|---|
Check out our online estimator overview at: https://www.sktime.org/en/stable/estimator_overview.html
**all_estimators()**
Signature:
all_estimators(
estimator_types=None,
filter_tags=None,
exclude_estimators=None,
return_names=True,
as_dataframe=False,
return_tags=None,
suppress_import_stdout=True,
)
Docstring:
Get a list of all estimators from sktime.
This function crawls the module and gets all classes that inherit
from sktime's and sklearn's base classes.
Not included are: the base classes themselves, classes defined in test
modules.from sktime.registry import all_estimators
estimators = all_estimators("forecaster", as_dataframe=True)
head_tail_horz(estimators.name, 5, 'Forecaster Estimators (some options)')
| Forecaster Estimators (some options) |
| name | |
|---|---|
| 0 | ARDL |
| 1 | ARIMA |
| 2 | AutoARIMA |
| 3 | AutoETS |
| 4 | AutoEnsembleForecaster |
| name | |
|---|---|
| 48 | UpdateEvery |
| 49 | UpdateRefitsEvery |
| 50 | VAR |
| 51 | VARMAX |
| 52 | VECM |
missing_data_estimators = all_estimators("forecaster",
filter_tags = {"handles-missing-data": True},
as_dataframe=True)
head_tail_horz(missing_data_estimators.name.sample(10), 5,
'Forecaster Estimators (using filters)')
| Forecaster Estimators (using filters) |
| name | |
|---|---|
| 6 | DirectTimeSeriesRegressionForecaster |
| 17 | TransformedTargetForecaster |
| 9 | ForecastingPipeline |
| 12 | NaiveForecaster |
| 11 | MultioutputTimeSeriesRegressionForecaster |
| name | |
|---|---|
| 10 | MultioutputTabularRegressionForecaster |
| 4 | DirRecTimeSeriesRegressionForecaster |
| 16 | StackingForecaster |
| 0 | ARIMA |
| 8 | ForecastByLevel |
| But can I not just use scikit-learn? |
|---|
In principle, yes, but many pitfalls ...
See our previous tutorial from the PyData Amsterdam 2020 for more details: https://github.com/sktime/sktime-tutorial-pydata-amsterdam-2020
Better: Use scikit-learn with sktime!
sktime provides a meta-estimator for this approach, which is:

| Using a Scikit-Learn Model |
|---|
| Airline Passenger Dataset (documentation) |
|---|
The classic Box & Jenkins airline data. Monthly totals of international airline passengers, 1949 to 1960.
Dimensionality: univariate Series length: 144 Frequency: Monthly Number of cases: 1
This data shows an increasing trend, non-constant (increasing) variance and periodic, seasonal patterns.
| Visualization: airline passenger dataset input |
|---|
from sktime.datasets import load_airline
from sktime.utils.plotting import plot_series
targets = load_airline()
plot_series(targets, colors = ['C6'], title = 'Airline Passenger Dataset');
train_targets, test_targets = temporal_train_test_split(targets, test_size=12)
horizon = ForecastingHorizon(test_targets.index, is_relative=False)
**`forecasting.compose.make_reduction()`**
Signature:
make_reduction(
estimator,
strategy='recursive',
window_length=10,
scitype='infer',
transformers=None,
pooling='local',
windows_identical=True,
)
Docstring:
Make forecaster based on reduction to tabular or time-series regression.
During fitting, a sliding-window approach is used to first transform the
time series into tabular or panel data, which is then used to fit a tabular or
time-series regression estimator. During prediction, the last available data is
used as input to the fitted regression estimator to generate forecasts.
Parameters
----------
estimator : an estimator instance
Either a tabular regressor from scikit-learn or a time series regressor from
sktime.
strategy : str, optional (default="recursive")
The strategy to generate forecasts. Must be one of "direct", "recursive" or
"multioutput".
window_length : int, optional (default=10)
Window length used in sliding window transformation.
scitype : str, optional (default="infer")
Legacy argument for downwards compatibility, should not be used.
`make_reduction` will automatically infer the correct type of `estimator`.
This internal inference can be force-overridden by the `scitype` argument.
Must be one of "infer", "tabular-regressor" or "time-series-regressor".
If the scitype cannot be inferred, this is a bug and should be reported.
transformers: list of transformers (default = None)
A suitable list of transformers that allows for using an en-bloc approach with
make_reduction. This means that instead of using the raw past observations of
y across the window length, suitable features will be generated directly from
the past raw observations. Currently only supports WindowSummarizer (or a list
of WindowSummarizers) to generate features e.g. the mean of the past 7
observations. Currently only works for RecursiveTimeSeriesRegressionForecaster.
pooling: str {"local", "global"}, optional
Specifies whether separate models will be fit at the level of each instance
(local) of if you wish to fit a single model to all instances ("global").
Currently only works for RecursiveTimeSeriesRegressionForecaster.
windows_identical: bool, (default = True)
Direct forecasting only.
Specifies whether all direct models use the same X windows from y (True: Number
of windows = total observations + 1 - window_length - maximum forecasting
horizon) or a different number of X windows depending on the forecasting horizon
(False: Number of windows = total observations + 1 - window_length
- forecasting horizon). See pictionary below for more information.| Visualization: SKLearn's KNeighbors with recursive strategy |
|---|
from sklearn.neighbors import KNeighborsRegressor
from sktime.forecasting.compose import make_reduction
regressor = KNeighborsRegressor(n_neighbors=2)
forecaster = make_reduction(regressor, strategy="recursive", window_length=15)
forecaster.fit(train_targets, fh=horizon)
predictions = forecaster.predict()
plot_series(train_targets, test_targets, predictions,
labels=["train_targets", "test_targets", "predictions"],
colors = ['C0', 'C1', 'C2'],
title = 'KNeighborsRegressor | strategy = "recursive"');
| Visualization: SKLearn's KNeighbors with multioutput strategy |
|---|
regressor = KNeighborsRegressor(n_neighbors=1)
forecaster = make_reduction(regressor, strategy="multioutput", window_length=7)
forecaster.fit(train_targets, fh=horizon)
predictions = forecaster.predict()
plot_series(train_targets, test_targets, predictions,
labels=["train_targets", "test_targets", "predictions"],
colors = ['C0', 'C1', 'C2'],
title = 'KNeighborsRegressor | strategy = "multioutput"');
| | Top | SciKit-Learn Way | SKTime Way | Multivariate | Panel Data | SKLearn & SKTime | Univariate Forecasting | Advanced Workflow | Forecasting with Exogeneous | Building a Forecaster | Time Series Classification | Time Series Regression | | ||
|---|---|---|
| More Advanced workflow |
| Data specification |
|---|
from sktime.forecasting.ets import AutoETS
data = load_airline()
plot_series(data, colors = ['C6'], title = 'Airline Data');
| Task specification |
|---|
# specifying the forecasting horizon: one year ahead, all months
# 12 steps ahead
horizon = np.arange(1, 13)
pretty(horizon, 'Forecast Horizon | np.arange(1, 13)')
| Forecast Horizon | np.arange(1, 13) |
| [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12] |
train_data = data.loc[:"1957-08"]
observed_data = train_data.copy()
observed_data.tail()
1957-04 348.00 1957-05 355.00 1957-06 422.00 1957-07 465.00 1957-08 467.00 Freq: M, Name: Number of airline passengers, dtype: float64
| Model specification |
|---|
forecaster = AutoETS(auto=True, sp=12, n_jobs=-1)
| Fitting |
|---|
forecaster.fit(train_data)
AutoETS(auto=True, n_jobs=-1, sp=12)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook.
AutoETS(auto=True, n_jobs=-1, sp=12)
| Prediction |
|---|
predictions = forecaster.predict(horizon)
plot_series(observed_data, predictions, colors = ['C2', 'C6'],
title = 'Predictions with AutoETS');
predictions
1957-09 413.46 1957-10 360.57 1957-11 314.50 1957-12 358.25 1958-01 363.38 1958-02 363.45 1958-03 417.74 1958-04 402.20 1958-05 398.85 1958-06 451.96 1958-07 498.86 1958-08 494.80 Freq: M, dtype: float64
| Observe New Data |
|---|
oberved_data = data.loc[:"1957-09"]
new_data = data.loc[["1957-09"]]
new_data
1957-09 404.00 Freq: M, Name: Number of airline passengers, dtype: float64
| Update |
|---|
forecaster.update(new_data)
/Users/evancarr/opt/anaconda3/envs/time_series_projects/lib/python3.10/site-packages/sktime/forecasting/base/_base.py:1881: UserWarning: NotImplementedWarning: AutoETS does not have a custom `update` method implemented. AutoETS will be refit each time `update` is called with update_params=True. To refit less often, use the wrappers in the forecasting.stream module, e.g., UpdateEvery. warn(
AutoETS(auto=True, n_jobs=-1, sp=12)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook.
AutoETS(auto=True, n_jobs=-1, sp=12)
| Predict again |
|---|
predictions = forecaster.predict(horizon)
plot_series(observed_data, predictions,
colors = ['C3', 'C4'],
title = 'Updated Model Predictions');
predictions
1957-10 354.74 1957-11 309.49 1957-12 352.61 1958-01 357.73 1958-02 357.87 1958-03 411.40 1958-04 396.16 1958-05 392.94 1958-06 445.34 1958-07 491.65 1958-08 487.74 1958-09 432.52 Freq: M, dtype: float64
| Understanding update() |
|---|
update_params = False. (True by default)forecaster.update?
Signature: forecaster.update(y, X=None, update_params=True) Docstring: Update cutoff value and, optionally, fitted parameters. If no estimator-specific update method has been implemented, default fall-back is as follows: update_params=True: fitting to all observed data so far update_params=False: updates cutoff and remembers data only State required: Requires state to be "fitted". Accesses in self: Fitted model attributes ending in "_". Pointers to seen data, self._y and self.X self.cutoff, self._is_fitted If update_params=True, model attributes ending in "_". Writes to self: Update self._y and self._X with `y` and `X`, by appending rows. Updates self.cutoff and self._cutoff to last index seen in `y`. If update_params=True, updates fitted model attributes ending in "_". Parameters ---------- y : time series in sktime compatible data container format Time series to which to fit the forecaster in the update. y can be in one of the following formats, must be same scitype as in fit: Series scitype: pd.Series, pd.DataFrame, or np.ndarray (1D or 2D) for vanilla forecasting, one time series Panel scitype: pd.DataFrame with 2-level row MultiIndex, 3D np.ndarray, list of Series pd.DataFrame, or nested pd.DataFrame for global or panel forecasting Hierarchical scitype: pd.DataFrame with 3 or more level row MultiIndex for hierarchical forecasting Number of columns admissible depend on the "scitype:y" tag: if self.get_tag("scitype:y")=="univariate": y must have a single column/variable if self.get_tag("scitype:y")=="multivariate": y must have 2 or more columns if self.get_tag("scitype:y")=="both": no restrictions on columns apply For further details: on usage, see forecasting tutorial examples/01_forecasting.ipynb on specification of formats, examples/AA_datatypes_and_datasets.ipynb X : time series in sktime compatible format, optional (default=None) Exogeneous time series to fit to Should be of same scitype (Series, Panel, or Hierarchical) as y if self.get_tag("X-y-must-have-same-index"), X.index must contain y.index there are no restrictions on number of columns (unlike for y) update_params : bool, optional (default=True) whether model parameters should be updated Returns ------- self : reference to self File: ~/opt/anaconda3/envs/time_series_projects/lib/python3.10/site-packages/sktime/forecasting/base/_base.py Type: method
| Automating the Process |
|---|
from sktime.forecasting.model_evaluation import evaluate
from sktime.forecasting.model_selection import ExpandingWindowSplitter
from sktime.utils.plotting import plot_windows
data = load_airline()
horizon = ForecastingHorizon(np.arange(12) + 1)
train_data, test_data = temporal_train_test_split(data, fh = horizon)
| Temporal Cross-Validation: ExpandingWindowSplitter() |
|---|
step_length = 3 - every new window will be three steps ahead[Cross-Validation Notebook](https://github.com/alan-turing-institute/sktime/blob/main/examples/window_splitters.ipynb) <- different window splitters and ways of doing temporal cross validation
cv = ExpandingWindowSplitter(step_length = 3,
fh = horizon,
initial_window = 10)
plot_windows(cv, data.iloc[:50])
| Backtesting: Evaluation using temporal cross-validaton |
|---|
forecaster = NaiveForecaster(strategy="last", sp=12)
cv = ExpandingWindowSplitter(step_length=12, fh=horizon, initial_window=72)
results = evaluate(forecaster=forecaster,
y=data,
cv=cv,
strategy="refit",
return_data=True)
results.iloc[:, :5].head()
| test_MeanAbsolutePercentageError | fit_time | pred_time | len_train_window | cutoff | |
|---|---|---|---|---|---|
| 0 | 0.16 | 0.00 | 0.00 | 72 | 1954-12 |
| 1 | 0.13 | 0.00 | 0.00 | 84 | 1955-12 |
| 2 | 0.11 | 0.00 | 0.00 | 96 | 1956-12 |
| 3 | 0.03 | 0.00 | 0.00 | 108 | 1957-12 |
| 4 | 0.11 | 0.00 | 0.00 | 120 | 1958-12 |
fig, ax = plot_series(
data,
results['y_pred'].iloc[0],
results['y_pred'].iloc[1],
results['y_pred'].iloc[2],
results['y_pred'].iloc[3],
results['y_pred'].iloc[4],
results['y_pred'].iloc[5],
labels=['y_pred'] + ["preds (Backtest " + str(x) + ")" for x in range(6)],
colors = ['C0', 'C1' 'C2', 'C3', 'C4', 'C5', 'C6']
)
ax.legend();
| Tuning |
|---|
| Advanced model building & composition |
|---|
window_length and n_neighbors simultaneously[9, 12, 15][1, 2, 3, 4, 5, 6, 7, 8, 9]from sktime.forecasting.model_selection import (ForecastingGridSearchCV,
SlidingWindowSplitter)
param_grid = {"window_length": [9, 12, 15],
"estimator__n_neighbors": np.arange(1, 10)}
regressor = KNeighborsRegressor()
forecaster = make_reduction(regressor,
strategy="recursive")
| SlidingWindowSplittler & ForcastingGridSearchCV |
|---|
**ForecastingGridSearchCV()**
Perform grid-search cross-validation to find optimal model parameters.
The forecaster is fit on the initial window and then temporal
cross-validation is used to find the optimal parameter.
Grid-search cross-validation is performed based on a cross-validation
iterator encoding the cross-validation scheme, the parameter grid to
search over, and (optionally) the evaluation metric for comparing model
performance. As in scikit-learn, tuning works through the common
hyper-parameter interface which allows to repeatedly fit and evaluate
the same forecaster with different hyper-parameters.
Parameters
----------
forecaster : estimator object
The estimator should implement the sktime or scikit-learn estimator
interface. Either the estimator must contain a "score" function,
or a scoring function must be passed.
cv : cross-validation generator or an iterable
e.g. SlidingWindowSplitter()
strategy : {"refit", "update", "no-update_params"}, optional, default="refit"
data ingestion strategy in fitting cv, passed to `evaluate` internally
defines the ingestion mode when the forecaster sees new data when window expands
"refit" = forecaster is refitted to each training window
"update" = forecaster is updated with training window data, in sequence provided
"no-update_params" = fit to first training window, re-used without fit or update
update_behaviour: str, optional, default = "full_refit"
one of {"full_refit", "inner_only", "no_update"}
behaviour of the forecaster when calling update
"full_refit" = both tuning parameters and inner estimator refit on all data seen
"inner_only" = tuning parameters are not re-tuned, inner estimator is updated
"no_update" = neither tuning parameters nor inner estimator are updated
param_grid : dict or list of dictionaries
Model tuning parameters of the forecaster to evaluate
scoring: function, optional (default=None)
Function to score models for evaluation of optimal parameters
n_jobs: int, optional (default=None)
Number of jobs to run in parallel.
None means 1 unless in a joblib.parallel_backend context.
-1 means using all processors.
refit: bool, optional (default=True)
True = refit the forecaster with the best parameters on the entire data in fit
False = best forecaster remains fitted on the last fold in cv
verbose: int, optional (default=0)
return_n_best_forecasters: int, default=1
In case the n best forecaster should be returned, this value can be set
and the n best forecasters will be assigned to n_best_forecasters_
pre_dispatch: str, optional (default='2*n_jobs')
error_score: numeric value or the str 'raise', optional (default=np.nan)
The test score returned when a forecaster fails to be fitted.
return_train_score: bool, optional (default=False)
backend: str, optional (default="loky")
Specify the parallelisation backend implementation in joblib, where
"loky" is used by default.
error_score : "raise" or numeric, default=np.nan
Value to assign to the score if an exception occurs in estimator fitting. If set
to "raise", the exception is raised. If a numeric value is given,
FitFailedWarning is raised.
Attributes
----------
best_index_ : int
best_score_: float
Score of the best model
best_params_ : dict
Best parameter values across the parameter grid
best_forecaster_ : estimator
Fitted estimator with the best parameters
cv_results_ : dict
Results from grid search cross validation
n_splits_: int
Number of splits in the data for cross validation
refit_time_ : float
Time (seconds) to refit the best forecaster
scorer_ : function
Function used to score model
n_best_forecasters_: list of tuples ("rank", <forecaster>)
The "rank" is in relation to best_forecaster_
n_best_scores_: list of float
The scores of n_best_forecasters_ sorted from best to worst
score of forecasterscv = SlidingWindowSplitter(window_length=60, fh=horizon)
gscv = ForecastingGridSearchCV(forecaster,
cv=cv,
param_grid=param_grid,
strategy="refit")
| Grid Search Cross-Validation - fit the training data with each set of parameters - predict with the forecasting horizon - create predictions generated from each combination of parameters - get the best evaluated combination of parameters |
|---|
gscv.fit(train_data)
ForecastingGridSearchCV(cv=SlidingWindowSplitter(fh=ForecastingHorizon([1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12], dtype='int64', is_relative=True),
window_length=60),
forecaster=RecursiveTabularRegressionForecaster(estimator=KNeighborsRegressor()),
param_grid={'estimator__n_neighbors': array([1, 2, 3, 4, 5, 6, 7, 8, 9]),
'window_length': [9, 12, 15]})In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. ForecastingGridSearchCV(cv=SlidingWindowSplitter(fh=ForecastingHorizon([1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12], dtype='int64', is_relative=True),
window_length=60),
forecaster=RecursiveTabularRegressionForecaster(estimator=KNeighborsRegressor()),
param_grid={'estimator__n_neighbors': array([1, 2, 3, 4, 5, 6, 7, 8, 9]),
'window_length': [9, 12, 15]})RecursiveTabularRegressionForecaster(estimator=KNeighborsRegressor())
KNeighborsRegressor()
KNeighborsRegressor()
predictions = gscv.predict(horizon)
plot_series(train_data, test_data, predictions,
labels = ['train_data', 'test_data', 'predictions'],
colors = ['C0', 'C1', 'C2'])
mean_absolute_percentage_error(predictions, test_data)
0.12411154270928935
| ForcastingGridSearchCV.best_params_ - returns best n_neighbors is 2 and window_length is 12 |
|---|
gscv.best_params_
{'estimator__n_neighbors': 2, 'window_length': 12}
plot_windows(cv, data.iloc[:84])
| Tuning and AutoML |
|---|
MultiplexForecaster - specify the selection of models / algorithms to be tested and comparedets - ExponentialSmoothing trend - from this articleExponential Smoothing Methods are a family of forecasting models. They use weighted averages of past observations to forecast new values. The idea is to give more importance to recent values in the series. Thus, as observations get older in time, the importance of these values get exponentially smaller.
from sktime.forecasting.compose import MultiplexForecaster
from sktime.forecasting.exp_smoothing import ExponentialSmoothing
from sktime.forecasting.naive import NaiveForecaster
forecaster = MultiplexForecaster(
forecasters=[
("naive", NaiveForecaster(strategy="last")),
("ets", ExponentialSmoothing(trend="add", sp=12)),],)
| Exponential Smoothing wins |
|---|
forecaster_param_grid = {"selected_forecaster": ["ets", "naive"]}
gscv = ForecastingGridSearchCV(forecaster, cv=cv, param_grid=forecaster_param_grid)
gscv.fit(train_data)
gscv.best_params_
{'selected_forecaster': 'ets'}
| Pipelining |
|---|
1) Transform the data 2) Fit the forecaster on the transformed data 3) During prediction, generate the forecast 4) Inverse transform the forecast to the original dimensions of the data
from sktime.forecasting.compose import TransformedTargetForecaster
from sktime.forecasting.trend import PolynomialTrendForecaster
from sktime.transformations.series.detrend import Deseasonalizer, Detrender
regressor = KNeighborsRegressor()
forecaster = make_reduction(regressor, strategy="recursive")
| Deseasonalizing, then detrending, then fitting the forecaster to the transformed data |
|---|
forecaster = TransformedTargetForecaster(
[("deseasonalize", Deseasonalizer(sp=12)),
("detrend", Detrender()),
("forecast", forecaster),])
| Pipeline then operates the same as any other forecaster, which can then fit and predict |
|---|
forecaster.fit(train_data)
predictions = forecaster.predict(horizon)
predictions
1960-01 397.97 1960-02 385.26 1960-03 428.56 1960-04 427.64 1960-05 443.90 1960-06 488.83 1960-07 526.62 1960-08 522.14 1960-09 486.26 1960-10 456.32 1960-11 409.21 1960-12 423.23 Freq: M, dtype: float64
| | Top | SciKit-Learn Way | SKTime Way | Multivariate | Panel Data | SKLearn & SKTime | Univariate Forecasting | Advanced Workflow | Forecasting with Exogeneous | Building a Forecaster | Time Series Classification | Time Series Regression | | ||
|---|---|---|
| Forecasting with Exogenous Variables |
This can be useful when there is one variable that is easier to make future predictions on that can then help predict other, more complicated variables
| Basic Workflow |
|---|
from sktime.datasets import load_longley
targets, inputs = load_longley()
targets.head()
Period 1947 60,323.00 1948 61,122.00 1949 60,171.00 1950 61,187.00 1951 63,221.00 Freq: A-DEC, Name: TOTEMP, dtype: float64
inputs.head()
| GNPDEFL | GNP | UNEMP | ARMED | POP | |
|---|---|---|---|---|---|
| Period | |||||
| 1947 | 83.00 | 234,289.00 | 2,356.00 | 1,590.00 | 107,608.00 |
| 1948 | 88.50 | 259,426.00 | 2,325.00 | 1,456.00 | 108,632.00 |
| 1949 | 88.20 | 258,054.00 | 3,682.00 | 1,616.00 | 109,773.00 |
| 1950 | 89.50 | 284,599.00 | 3,351.00 | 1,650.00 | 110,929.00 |
| 1951 | 96.20 | 328,975.00 | 2,099.00 | 3,099.00 | 112,075.00 |
horizon = np.arange(5) + 1
train_targets, test_targets, train_inputs, train_preds = temporal_train_test_split(targets, inputs, fh=horizon)
| - Exogenous data is passed as training data to the fit method - in predict, X is the exogenous data that corresponds to the forcasting horizon to generate the prediction |
|---|
forecaster = AutoARIMA()
forecaster.fit(train_targets, train_inputs)
predictions = forecaster.predict(horizon, X=train_preds)
plot_series(train_targets, test_targets, predictions,
labels=["train_targets", "test_targets", "predictions"],
colors = ['C1', 'C2', 'C3']);
| Pipelining with Exogeneous Data |
|---|
from sklearn.preprocessing import MinMaxScaler, PowerTransformer
from sktime.datasets import load_macroeconomic
from sktime.forecasting.compose import ForecastingPipeline
from sktime.transformations.series.adapt import TabularToSeriesAdaptor
from sktime.transformations.series.impute import Imputer
| Macroeconomic Dataset (documentation) |
|---|
US Macroeconomic Data for 1959Q1 - 2009Q3.
Dimensionality: multivariate, 14 Series length: 203 Frequency: Quarterly Number of cases: 1
This data is kindly wrapped via statsmodels.datasets.macrodata.
data = load_macroeconomic()
targets = data["unemp"]
inputs = data.drop(columns=["unemp"])
train_targets, test_targets, train_inputs, test_inputs = temporal_train_test_split(targets, inputs)
horizon = ForecastingHorizon(test_targets.index, is_relative=False)
| Processes in pipeline: - imputing missing data - scaling the data - the model for forecasting NOTE: transformers from many libraries can be used, as long as they are wrapped in the TabularToSeriesAdaptor wrapper When fit and predict are called, these transformations will be applied to the exogenous data rather than to the target data |
|---|
forecaster = ForecastingPipeline(
steps=[("imputer", Imputer(method="mean")),
("scale", TabularToSeriesAdaptor(MinMaxScaler(feature_range=(1, 2)))),
("boxcox", TabularToSeriesAdaptor(PowerTransformer(method="box-cox"))),
("forecaster", AutoARIMA(suppress_warnings=True)),])
forecaster.fit(y = train_targets, X = train_inputs)
predictions = forecaster.predict(fh = horizon, X = test_inputs)
plot_series(train_targets, predictions, test_targets,
labels=["train_targets", "predictions", "test_targets"],
colors = ['C4', 'C2', 'C1'])
(<Figure size 1600x400 with 1 Axes>, <AxesSubplot: ylabel='unemp'>)
| Multivariate Forecasting: Multiple Target Series |
|---|
all_estimators(
"forecaster",
filter_tags={"scitype:y": ["both", "multivariate"]},
return_names=False,
)
[sktime.forecasting.compose._column_ensemble.ColumnEnsembleForecaster, sktime.forecasting.dynamic_factor.DynamicFactor, sktime.forecasting.compose._ensemble.EnsembleForecaster, sktime.forecasting.compose._grouped.ForecastByLevel, sktime.forecasting.model_selection._tune.ForecastingGridSearchCV, sktime.forecasting.compose._pipeline.ForecastingPipeline, sktime.forecasting.model_selection._tune.ForecastingRandomizedSearchCV, sktime.forecasting.compose._multiplexer.MultiplexForecaster, sktime.forecasting.compose._pipeline.Permute, sktime.param_est.plugin.PluginParamsForecaster, sktime.forecasting.compose._pipeline.TransformedTargetForecaster, sktime.forecasting.var.VAR, sktime.forecasting.varmax.VARMAX, sktime.forecasting.vecm.VECM]
_, data = load_longley()
data = data.iloc[:, 2:4]
horizon = np.arange(3) + 1
training, testing = temporal_train_test_split(data, fh = horizon)
| The job is to predict both of the features at the same time |
|---|
training.head()
| UNEMP | ARMED | |
|---|---|---|
| Period | ||
| 1947 | 2,356.00 | 1,590.00 |
| 1948 | 2,325.00 | 1,456.00 |
| 1949 | 3,682.00 | 1,616.00 |
| 1950 | 3,351.00 | 1,650.00 |
| 1951 | 2,099.00 | 3,099.00 |
| By-Variable Ensembling |
|---|
from sktime.forecasting.compose import ColumnEnsembleForecaster
forecasters = [("trend", PolynomialTrendForecaster(), 0),
("ses", ExponentialSmoothing(), 1),]
forecaster = ColumnEnsembleForecaster(forecasters = forecasters)
forecaster.fit(training)
predictions = forecaster.predict(horizon)
predictions.head()
| UNEMP | ARMED | |
|---|---|---|
| 1960 | 3,688.65 | 2,552.43 |
| 1961 | 3,794.19 | 2,552.43 |
| 1962 | 3,899.72 | 2,552.43 |
| Bespoke multivariate models |
|---|
from sktime.forecasting.var import VAR
forecaster = VAR()
forecaster.fit(training)
predictions = forecaster.predict(horizon)
predictions.head()
| UNEMP | ARMED | |
|---|---|---|
| 1960 | 3,322.41 | 2,611.27 |
| 1961 | 3,153.43 | 2,673.11 |
| 1962 | 3,095.84 | 2,725.06 |
| | Top | SciKit-Learn Way | SKTime Way | Multivariate | Panel Data | SKLearn & SKTime | Univariate Forecasting | Advanced Workflow | Forecasting with Exogeneous | Building a Forecaster | Time Series Classification | Time Series Regression | | ||
|---|---|---|
| Building a Forecaster |
Check out our [forecasting extension template](https://github.com/alan-turing-institute/sktime/blob/main/extension_templates/forecasting.py)!
This is a Python file with to-do code blocks that allow you to implement your own, sktime-compatible forecasting algorithm.
| Summary |
|---|
| Useful Resources |
|---|
| | Top | SciKit-Learn Way | SKTime Way | Multivariate | Panel Data | SKLearn & SKTime | Univariate Forecasting | Advanced Workflow | Forecasting with Exogeneous | Building a Forecaster | Time Series Classification | Time Series Regression | | ||
|---|---|---|
| Time Series Classification |
| Univariate time series classification |
|---|
In univariate time series classification, we have a single time series variable and an associated label for multiple instances. The goal is to find a classifier that can learn the relationship between the time series and labels, and accurately predict the label of a new, unlabelled series. sktime provides time series classification algorithms and tools for building composite machine learning models.
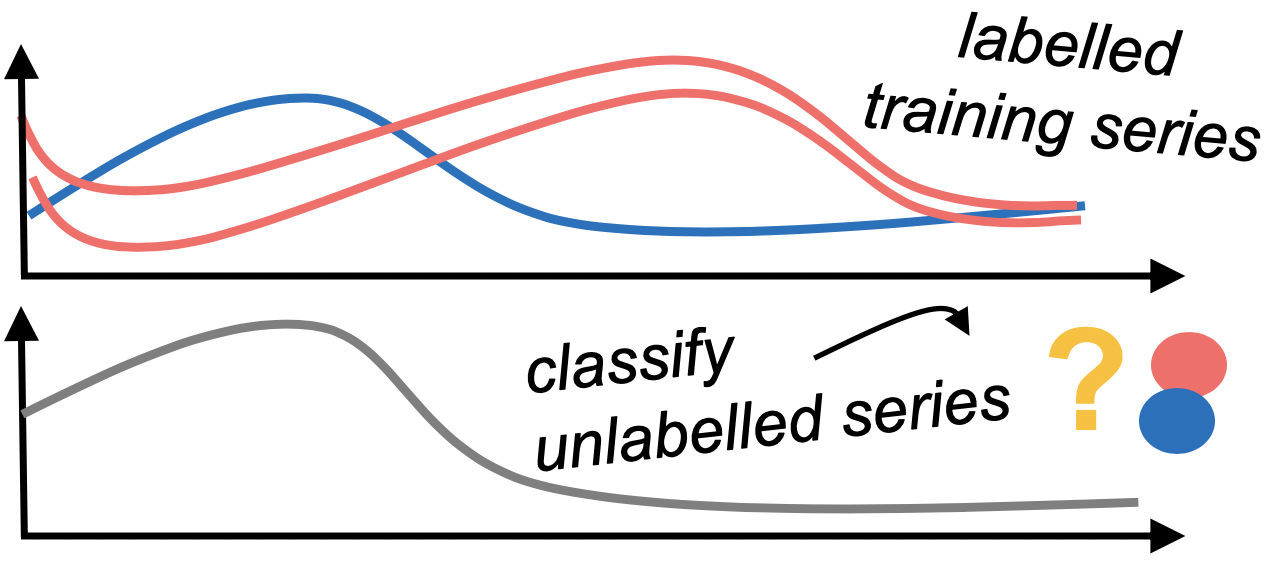
| The basic workflow |
|---|
| Data |
|---|
To find other datasets, go to: http://timeseriesclassification.com/dataset.php
from warnings import simplefilter
simplefilter(action="ignore", category=FutureWarning)
| UCR UEA Dataset (documentation) |
|---|
Load dataset from UCR UEA time series archive.
Downloads and extracts dataset if not already downloaded. Data is assumed to be in the standard .ts format: each row is a (possibly multivariate) time series. Each dimension is separated by a colon, each value in a series is comma separated. For examples see sktime.datasets.data.tsc. ArrowHead is an example of a univariate equal length problem, BasicMotions an equal length multivariate problem.
import matplotlib.pyplot as plt
import numpy as np
from sktime.datasets import load_UCR_UEA_dataset
from sktime.datatypes import convert
inputs, targets = load_UCR_UEA_dataset("ItalyPowerDemand", return_X_y=True)
see(inputs.head(3), 'Inputs before converting to NumPy 3D')
| Inputs before converting to NumPy 3D |
| dim_0 | |
|---|---|
| 0 | 0 -0.71 1 -1.18 2 -1.37 3 -1.59 4 -1.47 5 -1.37 6 -1.09 7 0.05 8 0.93 9 1.09 10 1.28 11 0.96 12 0.61 13 0.01 14 -0.65 15 -0.27 16 -0.21 17 0.61 18 1.37 19 1.46 20 1.05 21 0.58 22 0.17 23 -0.27 dtype: float64 |
| 1 | 0 -0.99 1 -1.43 2 -1.58 3 -1.61 4 -1.63 5 -1.38 6 -1.02 7 -0.36 8 0.72 9 1.20 10 1.12 11 1.05 12 0.79 13 0.46 14 0.49 15 0.56 16 0.61 17 0.31 18 0.26 19 1.10 20 1.05 21 0.69 22 -0.05 23 -0.38 dtype: float64 |
| 2 | 0 1.32 1 0.57 2 0.20 3 -0.09 4 -0.18 5 -0.27 6 -0.09 7 -1.40 8 -1.12 9 -0.74 10 0.01 11 -0.09 12 0.01 13 -0.46 14 -0.55 15 -0.74 16 -0.74 17 -0.74 18 -1.12 19 -0.46 20 0.48 21 2.35 22 2.26 23 1.60 dtype: float64 |
inputs = convert(inputs, from_type="nested_univ", to_type="numpy3D")
| This data: - 1,096 different individuals' records - 1 variable - for 24 time points |
|---|
pretty(inputs.shape, 'inputs.shape')
| inputs.shape |
| (1096, 1, 24) |
# binary target variable
pretty(np.unique(targets), 'np.unique(targets)')
| np.unique(targets) |
| ['1', '2'] |
labels, counts = np.unique(targets, return_counts=True)
fig, ax = plt.subplots(1, figsize=plt.figaspect(0.25))
for label in labels:
ax.plot(inputs[targets == label, 0, :][0], label=f"class {label}")
ax.set(ylabel="Scaled distance from midpoint", xlabel="Index");
| Train-test split |
|---|
from sklearn.model_selection import train_test_split
train_in, test_in, train_out, test_out = train_test_split(inputs, targets)
| Model specification |
|---|
Find out more about ROCKET:
from sktime.classification.kernel_based import RocketClassifier
classifier = RocketClassifier()
| Fitting |
|---|
%%time
classifier.fit(train_in, train_out)
RocketClassifier()In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook.
RocketClassifier()
| The X here is panel data with time index data for multiple features. All SKTime fit methods need this format for their X input. |
|---|
classifier.fit?
Signature: classifier.fit(X, y) Docstring: Fit time series classifier to training data. Parameters ---------- X : 3D np.array (any number of dimensions, equal length series) of shape [n_instances, n_dimensions, series_length] or 2D np.array (univariate, equal length series) of shape [n_instances, series_length] or pd.DataFrame with each column a dimension, each cell a pd.Series (any number of dimensions, equal or unequal length series) or of any other supported Panel mtype for list of mtypes, see datatypes.SCITYPE_REGISTER for specifications, see examples/AA_datatypes_and_datasets.ipynb y : 1D np.array of int, of shape [n_instances] - class labels for fitting indices correspond to instance indices in X Returns ------- self : Reference to self. Notes ----- Changes state by creating a fitted model that updates attributes ending in "_" and sets is_fitted flag to True. File: ~/opt/anaconda3/envs/time_series_projects/lib/python3.10/site-packages/sktime/classification/base.py Type: method
| Prediction |
|---|
predictions = classifier.predict(test_in)
| Evaluation |
|---|
from sklearn.metrics import accuracy_score
accuracy_score(test_out, predictions)
0.9708029197080292
| Classifiers in SKTIme |
|---|
from sktime.registry import all_estimators
all_estimators("classifier", return_names=False)
[sktime.classification.kernel_based._arsenal.Arsenal, sktime.classification.dictionary_based._boss.BOSSEnsemble, sktime.classification.deep_learning.cnn.CNNClassifier, sktime.classification.interval_based._cif.CanonicalIntervalForest, sktime.classification.feature_based._catch22_classifier.Catch22Classifier, sktime.classification.compose._pipeline.ClassifierPipeline, sktime.classification.compose._column_ensemble.ColumnEnsembleClassifier, sktime.classification.compose._ensemble.ComposableTimeSeriesForestClassifier, sktime.classification.dictionary_based._cboss.ContractableBOSS, sktime.classification.interval_based._drcif.DrCIF, sktime.classification.dummy._dummy.DummyClassifier, sktime.classification.distance_based._elastic_ensemble.ElasticEnsemble, sktime.classification.deep_learning.fcn.FCNClassifier, sktime.classification.feature_based._fresh_prince.FreshPRINCE, sktime.classification.hybrid._hivecote_v1.HIVECOTEV1, sktime.classification.hybrid._hivecote_v2.HIVECOTEV2, sktime.classification.dictionary_based._boss.IndividualBOSS, sktime.classification.dictionary_based._tde.IndividualTDE, sktime.classification.distance_based._time_series_neighbors.KNeighborsTimeSeriesClassifier, sktime.classification.deep_learning.lstmfcn.LSTMFCNClassifier, sktime.classification.deep_learning.mlp.MLPClassifier, sktime.classification.dictionary_based._muse.MUSE, sktime.classification.feature_based._matrix_profile_classifier.MatrixProfileClassifier, sktime.classification.early_classification._probability_threshold.ProbabilityThresholdEarlyClassifier, sktime.classification.distance_based._proximity_forest.ProximityForest, sktime.classification.distance_based._proximity_forest.ProximityStump, sktime.classification.distance_based._proximity_forest.ProximityTree, sktime.classification.feature_based._random_interval_classifier.RandomIntervalClassifier, sktime.classification.interval_based._rise.RandomIntervalSpectralEnsemble, sktime.classification.deep_learning.resnet.ResNetClassifier, sktime.classification.kernel_based._rocket_classifier.RocketClassifier, sktime.classification.distance_based._shape_dtw.ShapeDTW, sktime.classification.shapelet_based._stc.ShapeletTransformClassifier, sktime.classification.feature_based._signature_classifier.SignatureClassifier, sktime.classification.compose._pipeline.SklearnClassifierPipeline, sktime.classification.feature_based._summary_classifier.SummaryClassifier, sktime.classification.interval_based._stsf.SupervisedTimeSeriesForest, sktime.classification.feature_based._tsfresh_classifier.TSFreshClassifier, sktime.classification.deep_learning.tapnet.TapNetClassifier, sktime.classification.dictionary_based._tde.TemporalDictionaryEnsemble, sktime.classification.interval_based._tsf.TimeSeriesForestClassifier, sktime.classification.kernel_based._svc.TimeSeriesSVC, sktime.classification.dictionary_based._weasel.WEASEL, sktime.classification.compose._ensemble.WeightedEnsembleClassifier]
| But can I not just use scikit-learn? |
|---|
In principle, yes, but may not be as powerful as dedicated time series classification algorithms ...
See our previous tutorial from the PyData Amsterdam 2020 for more details: https://github.com/sktime/sktime-tutorial-pydata-amsterdam-2020
Compare algorithms from sktime and scikit-learn!
from sklearn.neighbors import KNeighborsClassifier
from sklearn.pipeline import make_pipeline
from sktime.transformations.panel.reduce import Tabularizer
| Tabularizer() converts panel data to cross-sectional data - this makes it possible to use other classifiers such as SKLearn models with the data - previously the data had been formatted for use with SKTime specifically - The difference in classifiers is that SKTime's classifiers are dedicated time series classifiers rather than just generic models for classification |
|---|
classifier = make_pipeline(Tabularizer(),
KNeighborsClassifier(n_neighbors=1,
metric="euclidean"))
classifier.fit(train_in, train_out)
Pipeline(steps=[('tabularizer', Tabularizer()),
('kneighborsclassifier',
KNeighborsClassifier(metric='euclidean', n_neighbors=1))])In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. Pipeline(steps=[('tabularizer', Tabularizer()),
('kneighborsclassifier',
KNeighborsClassifier(metric='euclidean', n_neighbors=1))])Tabularizer()
KNeighborsClassifier(metric='euclidean', n_neighbors=1)
predictions = classifier.predict(test_in)
accuracy_score(test_out, predictions)
0.9817518248175182
| Advanced Model Building and Composition |
|---|
| Pipelining |
|---|
Check out the tsfresh package for automatic feature extraction: https://tsfresh.readthedocs.io/en/latest/
from sklearn.ensemble import RandomForestClassifier
from sklearn.pipeline import make_pipeline
from sktime.transformations.panel.tsfresh import TSFreshFeatureExtractor
# %%time
# classifier = make_pipeline(TSFreshFeatureExtractor(disable_progressbar=True,
# show_warnings=False),
# RandomForestClassifier(),)
# classifier.fit(train_in, train_out)
# predictions = classifier.predict(test_in)
# accuracy_score(test_out, predictions)
| | Top | SciKit-Learn Way | SKTime Way | Multivariate | Panel Data | SKLearn & SKTime | Univariate Forecasting | Advanced Workflow | Forecasting with Exogeneous | Building a Forecaster | Time Series Classification | Time Series Regression | | ||
|---|---|---|
| Time Series Regression |
| The basic workflow |
|---|
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sktime.datasets import load_UCR_UEA_dataset
Find out more about the dataset here: https://zenodo.org/record/3902673#.YXqxNy8w3UI
X, y = load_UCR_UEA_dataset(name="ChlorineConcentration", return_X_y=True)
print(X.shape, y.shape)
X_train, X_test, y_train, y_test = train_test_split(X, y)
fig, ax = plt.subplots(1)
ax.hist(y)
ax.set(xlabel="target variable (bins)", ylabel="frequency");
# fig, ax = plt.subplots(1, figsize=plt.figaspect(0.25))
# for i in range(5):
# ax.plot(X_train)
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import mean_squared_error
from sklearn.pipeline import make_pipeline
from sktime.transformations.panel.rocket import Rocket
regressor = make_pipeline(Rocket(), RandomForestRegressor())
%%time
regressor.fit(train_in, train_out)
Pipeline(steps=[('rocket', Rocket()),
('randomforestregressor', RandomForestRegressor())])In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. Pipeline(steps=[('rocket', Rocket()),
('randomforestregressor', RandomForestRegressor())])Rocket()
RandomForestRegressor()
# predictions = regressor.predict(test_in)
# mean_squared_error(test_out, predictions)
| Reducing forecasting to time series |
|---|

import numpy as np
from sktime.datasets import load_airline
from sktime.forecasting.compose import make_reduction
from sktime.forecasting.model_selection import temporal_train_test_split
from sktime.utils.plotting import plot_series
data = load_airline()
horizon = np.arange(12) + 1
training, testing = temporal_train_test_split(data, fh=horizon)
| Must specify that this is a time-series regressor rather than a tabular regressor, i.e. the scitype |
|---|
forecaster = make_reduction(
regressor, scitype="time-series-regressor", window_length=12)
| Look up the term "scitype" in our glossary: |
|---|
https://www.sktime.org/en/stable/glossary.html#term-Scientific-type
from sktime.forecasting.compose import TransformedTargetForecaster
from sktime.transformations.series.detrend import Detrender
pipe = TransformedTargetForecaster([("detrend",
Detrender()), ("forecast",
forecaster)])
%%time
pipe.fit(training)
TransformedTargetForecaster(steps=[('detrend', Detrender()),
('forecast',
RecursiveTimeSeriesRegressionForecaster(estimator=Pipeline(steps=[('rocket',
Rocket()),
('randomforestregressor',
RandomForestRegressor())]),
window_length=12))])In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. TransformedTargetForecaster(steps=[('detrend', Detrender()),
('forecast',
RecursiveTimeSeriesRegressionForecaster(estimator=Pipeline(steps=[('rocket',
Rocket()),
('randomforestregressor',
RandomForestRegressor())]),
window_length=12))])predictions = pipe.predict(horizon)
plot_series(training, testing, predictions,
labels=["y_train", "y_test", "y_pred"],
colors = ['C0', 'C1', 'C2']);
| Multivariate Time Series Classification |
|---|
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.pipeline import Pipeline
from sktime.classification.compose import ColumnEnsembleClassifier
from sktime.classification.dictionary_based import BOSSEnsemble
from sktime.classification.interval_based import TimeSeriesForestClassifier
from sktime.transformations.panel.compose import ColumnConcatenator
| Loading Multivariate Time Series/Panel Data |
|---|
The data set we use in this notebook was generated as part of a student project where four students performed four activities whilst wearing a smart watch. The watch collects 3D accelerometer and a 3D gyroscope It consists of four classes, which are walking, resting, running and badminton. Participants were required to record motion a total of five times, and the data is sampled once every tenth of a second, for a ten second period.
input_data, target_data = load_UCR_UEA_dataset("BasicMotions", return_X_y=True)
input_data = convert(input_data, from_type="nested_univ", to_type="numpy3D")
train_in, test_in, train_out, test_out = train_test_split(input_data, target_data)
pretty(train_in.shape, 'train_in.shape')
pretty(train_out.shape, 'train_out.shape')
pretty(test_in.shape, 'test_in.shape')
pretty(test_out.shape, 'test_out.shape')
| train_in.shape |
| (60, 6, 100) |
| train_out.shape |
| (60,) |
| test_in.shape |
| (20, 6, 100) |
| test_out.shape |
| (20,) |
# multi-class target variable
np.unique(train_out)
array(['badminton', 'running', 'standing', 'walking'], dtype='<U9')
| Multivariate Classification |
|---|
sktime offers three main ways of solving multivariate time series classification problems:
ColumnConcatenator and apply a classifier to the concatenated data,ColumnEnsembleClassifier in which one classifier is fitted for each time series column and their predictions aggregated,| Time Series Concatenation |
|---|
We can concatenate multivariate time series/panel data into long univiariate time series/panel and then apply a classifier to the univariate data.
steps = [
("concatenate", ColumnConcatenator()),
("classify", TimeSeriesForestClassifier(n_estimators=100)),]
classifier = Pipeline(steps)
classifier.fit(train_in, train_out)
classifier.score(test_in, test_out)
1.0
| Column Ensembling |
|---|
We can also fit one classifier for each time series column and then aggregated their predictions. The interface is similar to the familiar ColumnTransformer from sklearn.
classifier = ColumnEnsembleClassifier(
estimators=[
("TSF0", TimeSeriesForestClassifier(n_estimators=10), [0]),
("BOSSEnsemble3", BOSSEnsemble(max_ensemble_size=5), [3]),])
# classifier.fit(X_train, y_train)
classifier.score(X_test, y_test)
| Bespoke classification algorithms |
|---|
Another approach is to use bespoke (or classifier-specific) methods for multivariate time series data. Here, we try out the HIVE-COTE (version 2) algorithm in multidimensional space.
Check out the research paper: https://link.springer.com/article/10.1007%2Fs10994-021-06057-9
from sktime.classification.hybrid import HIVECOTEV2
X_train, y_train = load_UCR_UEA_dataset("BasicMotions", split="train", return_X_y=True)
X_test, y_test = load_UCR_UEA_dataset("BasicMotions", split="test", return_X_y=True)
| HiveCoteV2 is current time series state of the art |
|---|
classifier = HIVECOTEV2(
stc_params={"n_shapelet_samples": 1000},
drcif_params={"n_estimators": 25},
arsenal_params={"n_estimators": 10},
tde_params={"n_parameter_samples": 100},
verbose=0,)
%%time
classifier.fit(train_in, train_out)
HIVECOTEV2(arsenal_params={'n_estimators': 10},
drcif_params={'n_estimators': 25},
stc_params={'n_shapelet_samples': 1000},
tde_params={'n_parameter_samples': 100})In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. HIVECOTEV2(arsenal_params={'n_estimators': 10},
drcif_params={'n_estimators': 25},
stc_params={'n_shapelet_samples': 1000},
tde_params={'n_parameter_samples': 100})predictions = classifier.predict(test_in)
accuracy_score(test_out, predictions)
0.95
| Building a Classifier |
|---|
Check out our classifier extension template!
This is a Python file with to-do code blocks that allow you to implement your own, sktime-compatible classification algorithm.
| Summary |
|---|
| Useful Resources |
|---|
| Deep Learning |
|---|